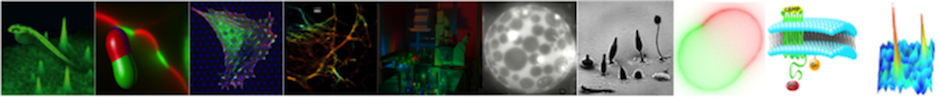
Recent experimental evidence suggests that the nanostructure of cell membranes is of crucial importance for a plethora of biological functions. The transduction of signals from the outside to the inside of the cell, for example, is mediated via membrane bound receptor proteins. Because receptors are the first element in a signaling cascade that can result in a multitude of cellular responses, they are currently the most important drug target. Receptors are believed to fulfill their signaling task with the help of, and in interaction with nanometer sized membrane domains. Therefore, an increased fundamental understanding of the nanoscopic membrane organization is believed to pave the way for more efficient drug design and improved drug quality.
Recent experimental evidence suggests that the nanostructure of cell membranes is of crucial importance for a plethora of biological functions. The transduction of signals from the outside to the inside of the cell, for example, is mediated via membrane bound receptor proteins. Because receptors are the first element in a signaling cascade that can result in a multitude of cellular responses, they are currently the most important drug target. Receptors are believed to fulfill their signaling task with the help of, and in interaction with nanometer sized membrane domains. Therefore, an increased fundamental understanding of the nanoscopic membrane organization is believed to pave the way for more efficient drug design and improved drug quality.
In addition, membranes are a supreme example for a very complex and multifunctional self-assembling nanosystem. They might therefore serve as a paradigm for developing bionic applications from the bottom up. One of the many examples is the use of membrane receptors adsorbed on substrates as biosensors in lab-on-a-chip applications.
Given the complexity of a living cell, all observations and subsequent models leave room for ambiguity in interpretation. We set out to develop and study model systems of biological membranes in order to drastically restrict the number of (un-determined) molecular players and for rigorously tests of current theoretical models. We use artificial membrane model systems, giant unilamellar vesicles (GUVs) and supported lipid bilayers, to study lipid heterogeneities, domain formation and stability, and the effects of mechanical forces on pure, mixed and coated biomembranes.
GUVs mimicking the lipid composition of the plasma membrane of living cells exhibit separation and coexistence of micron sized liquid-ordered and liquid-disordered phases below a critical temperature. By accurate determination of the membrane material parameters, we could show that in living cells lipid domains must have a size not larger than 10 nm.
GUVs are also excellent model systems in order to learn about the processes and the mechanical basis for the formation of complex intracellular membrane networks like the endoplasmic reticulum. GUVs coated with microtubule-associated motors (kinesin, NCD), when plated onto microtubule-coated support, spontaneously form hollow, nanometer-sized membrane tubes. This process resembles that of the cell which builds up dynamic internal structures. Here we are interested in the interplay of – the mechanical properties of the membrane, the concentration and type (processive, non-processive) of the motors, the availability of microtubule tracks – on the size, shape and dynamics of the final membrane nanotube network.
in collaboration with:
Marileen Dogterom, AMOLF
Cees Storm, TU Eindhoven
latest results:
Tube regulation by diffusion: Paige Shaklee, Timon Idema, Marileen Dogterom, Cees Storm & Thomas Schmidt. “Kinesin recycling in stationary membrane tubes.”, Biophys. J. (2010) 99:1835.
GUVs as drug-screen: Stefan Semrau, Marilyn Monster, Matthijs van der Knaap, Bobby Florea, Thomas Schmidt & Mark Overhand. “Membrane lysis by gramicidin S visualized in red blood cells and giant vesicles.”, BBA Biomembranes (2010) 1798:2033